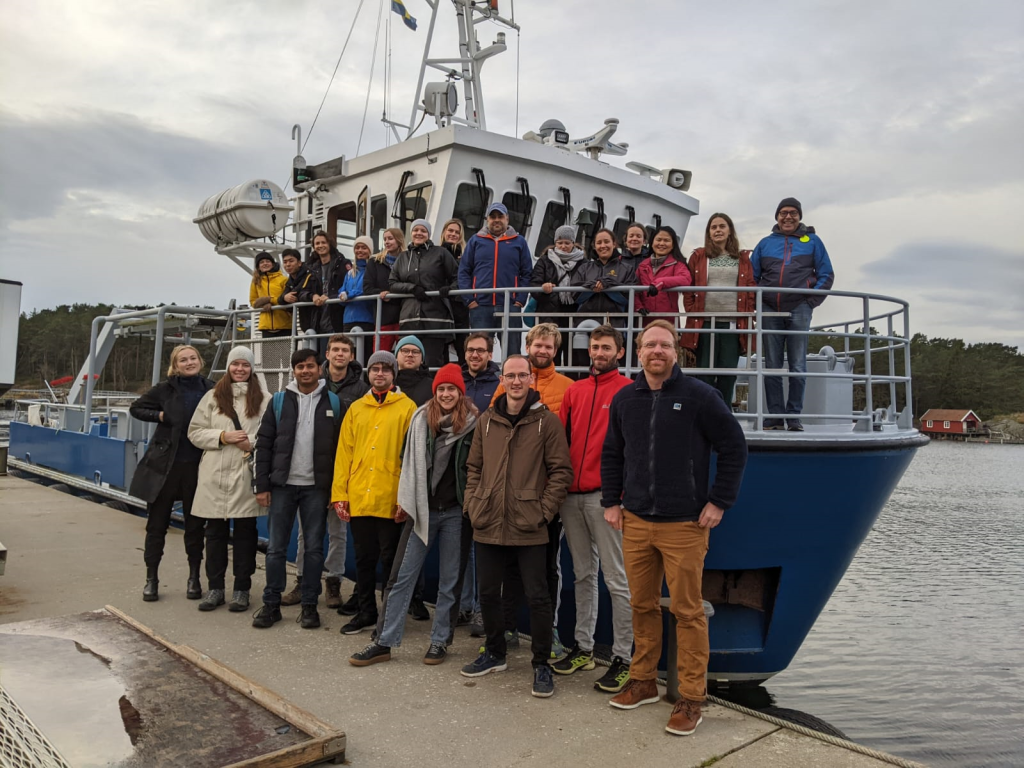
The 6th course on “eScience tools in climate science: linking observations with modelling” was held from 31.10.-11.11.2022 in Tjärnö, Marine Laboratory, Sweden with 17 students from Sweden, Norway, Denmark, Finland, Canada, South Africa and Australia. The course was held with support from the Universities of Stockholm and Oslo, the EU project CRICES, the Norwegian research school CHESS as well as the Bolin Center and the Norwegian Meteorological Institute. Advanced computing resources were provided by the CESNET resource centre, co-funded by the EGI-ACI project. Training material and IT infrastructure was for its “cloud” part prepared by the Pangeo Europe Team, in particular Guillaume Eynard-Bontemps and Jean Iaquinta. Anne Fouilloux and Tina Odaka. Data were obtained from the pangeo hub, cmip6 and ACTRIS. Further satellite, reanalysis and model data were assembled by assistants and Jan Griesfeller in the first days of the course and stored “read-only” on the s3 store at the Norwegian Meteorological Institute.
Students were working in small groups, assisted by 7 highly motivated and engaged assistants. Each student had to work through a small study within a science topic, proposed by the assistant. Topics dealt with were biological production in the Arctic ocean in a world with less sea ice along with changes in ocean salinity, phytoplancton blooming and DMS emissions. Atmospheric processes studied were aerosol size distribution variability, aerosol removal by precipitation, aerosol transport along with atmospheric rivers, but also ozone changes in the stratosphere after volcanic eruptions. Engagement and cross-talk during the course was great. Introductions to the Marine laboratory, a research vessl trip to Koster with its marine biology museum and a suite of lectures accompanied the group work.
With some hickups, all students and assistants managed to login to the EOSC Pangeo JupyterHub within the first days. Usage of the jupyterhub throughout the course was working very well and no crashes where observed. Training material served its purpose very well and the students were able to learn from example notebooks. Groups stored their own jupyter notebooks and reports in a github repository, based on a template provided for the course. Github integration into the jupyterhub was highly appreciated. Examples were added to the tutorial during the course. The discussion section in the overall course github repository was lively used to exchange tips and tricks. The EOSC Pangeo storage was unfortunately too complicated to establish and use. Only 2 assistants managed to login in, probably also because of limited time to test this further. Additional data were assembled and provided read only at the METNO s3 store with external help, which was more convenient for the students and assistants. However, reading data from this s3 store became slow when all students started to analyse and load larger data sets in the second week of the course. Students certainly got a glimpse of problems and solutions when dealing with large datasets. This part could be probably taught and instructed a bit better. Altogether all students finalised a report draft, participated in an internal review process and presented their findings in a hybrid meeting two weeks after the course.
Text and photo: Michael Schulz